Listen to this record
Letting Claude Loose in The Wet Lab
Giving Claude access to the wet lab with ResearchOS.
Origins
The moment of inspiration that became Tetsuwan came on November 6th of 2023, during OpenAI’s first DevDay. I was sitting around a TV with 15 other people in an office in Manhattan, watching the reveal of GPT-4o. It felt strange and unfamiliar to watch something like this with others, as if we were all watching the beginning of a change that none of us could really comprehend. That feeling of strangeness has returned to me this week, as we release our first computer use demo for lab automation.
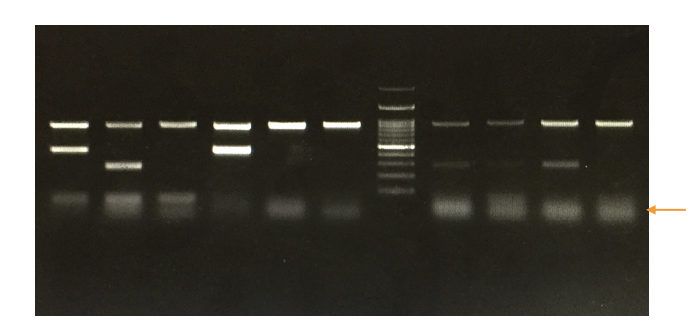
The first thing I did after that stream was run over to my laptop to throw a picture of a gel at GPT-4o (what this looks like is pictured above). The purpose of a (electrophoresis) gel is to size a fragment of DNA, a common troubleshooting/QC step in molecular biology workflows. Those dark ovals at the top are where fragments are pipetted in, and then an electric gradient is applied across the gel. The bigger fragments drag more in the gel and move slower, while the smaller fragments move faster. The first and last lanes contain “ladder”, a set of standard sized fragments that allow the researcher to size the band against the standard. As you can see, gels can be pretty blurry. Those faint bands below the brighter ones are called “primer dimers”. They are artifacts that result from the non-specific amplification of DNA (the polymerase, the chemical copy machine used in a polymerase chain reaction, puts more the wrong things in the copy machine).
The task that I handed 4o was to size the bands relative to the ladder. Of course, just because the tool might be capable, it is not necessarily the right tool (1). But to my surprise, not only was the model capable of properly sizing the bands, but it had caught a primer dimer that I did not immediately notice. It told me to change my annealing temperature to get rid of it. I felt like I had just seen a magic trick (2). It became clear at that moment that one day these models would be married with lab robots to provide them with physical agency, and that the process of research would begin to significantly change. Tetsuwan has been chasing that change since then.
Karnofsky’s Duplicator
In our first blog post back in 2024, I wrote about Holden Karnofsky's idea of "the duplicator." The duplicator is a thought experiment (borrowed from Calvin & Hobbes) in which humanity invents a device capable of instantly cloning any person, effectively scaling the human population of innovators without growing the broader population. The larger idea, central to Karnofsky's concept of a Process to Automate Scientific and Technological Advancement (PASTA), is that the most radical invention imaginable would be something that removes the fundamental bottleneck on progress: the scarcity of human minds. Karnofsky writes:
I think the Duplicator would be a more powerful technology than warp drives, tricorders, laser guns, or even teleporters. Minds are the source of innovation that can lead to all of those other things. So being cheaply able to duplicate them would be an extraordinary situation. - Holden Karnofsky, in "The Duplicator”
I followed this quote by writing:
“We cannot physically clone the minds of existing human scientists, but large language models are becoming increasingly capable of completing many of the cognitive tasks that underlie the scientific method, including hypothesis generation, experiment execution & troubleshooting, and knowledge distillation. These developments have occurred at breakneck speed. However, these models are limited in silico, especially for discovery in the physical sciences. Unlike human scientists, they cannot interact with the world and experiment to test and refine their hypotheses. Biology will not be solved in silico soon; rather, it will require experimentation and the collection of massive amounts of reproducible data to enable more robust simulations. Combining language models with lab robotics provides them with the agency to physically interact with the world to progress the scientific method similar to human scientists. The vision of the duplicator and unlimited scientific talent will be achieved through combining these models with physical agency, enabling them to interact and experiment in the physical world.”
The Problem
Gingko & OpenAI put out a very cool paper recently on marrying their automation hardware platform with GPT to optimize a cell-free protein expression (CFPE) chemistry (3). This is the first non-trivial example of what an embodied language model could look like and accomplish in the wet lab, and has only stoked the flames of a red-hot space. While we are excited about this work, we were not surprised to hear about some of the challenges the team faced. The Ginkgo-OpenAI team shared at SLAS this year that the model would try to do funny things like aspirate negative volume out of containers or pipette nanoliters instead of microliters, leaving wells virtually unchanged. Models are capable of doing really cool things, but the process of lab automation is still something that they struggle with, without well (and meticulously) engineered systems that augment their capabilities.
As highlighted in previous blog posts, the ability to stand up and iterate on automated workflows is not a new challenge for embodiment in the lab. We’ve got a ridiculous number of lab robots of varying form factors and functionalities. We’ve got models capable of generating hypotheses and entire apps. Why is it so difficult to “just put 2 and 2 together” here (4)? Long story short, it’s difficult for researchers, lab robots, and models to all speak the same language. As Abhishaike Morijan writes: “Most lab protocols can be automated, they just often aren’t worth automating”.
Why aren’t most protocols worth automating? Because the process of translating the way that we as human researchers understand our experiments at (workflows riddled with assumptions and tacit knowledge) into an explicit set of instructions is called automation engineering and it is not easy (5). Investing the amount of time and effort this process of translation requires only makes sense when you are executing a single workflow a LOT of times. If you ask automation engineers what they actually spend their time doing, many will tell you that writing the script is the simple part of this translation, and that they only spend ~20% of their time coding (6). This is less of a software engineering/writing code challenge and more of a description problem, hence why approaches that treat it as a normal code generation problem struggle.
It seldom makes sense to invest the days, weeks, or even months needed to move a workflow from the bench to automation. The only time this activation energy makes sense to pay is when a workflow is iterated thousands (or hundreds of thousands) of times, with little variation. These workflows are referred to as “low-mix, high-throughput” and resemble a process that would be performed at an assembly line more than one that would be done at a lab bench. Without solving this process, every new experiment a researcher (or model) might want to put onto an automated platform would require a prohibitive amount of time and effort. This is not an immediately obvious obstacle, especially to those who have not yet been subject to lab automation, but it is a massive one.
So, we have spent the last 2 years iterating on solutions to this answer. You can read more about it here. The summary of the product is that it uses a model, armed with our two domain-specific languages (DSLs) to structure a script ingested by our own compiler and ultimately turned into automation code. It structures the gap between implicit and explicit, so that researchers (implicit beings), robots (explicit things), and models (explicit things that are good at communicating implicitly) can seamlessly collaborate. The blog linked in this paragraph talks more about this, but you can also check out OwlPosting and Scaling Biotech’s blogs on the topic. While our work at Tetsuwan revolves more around making the glue between all of these pieces (researchers, robots, and models) rather than using a combination of them to solve a specific question, we figured it would be cool to show off the type of collaboration ResearchOS enables.
Computer-Use with Claude & ResearchOS
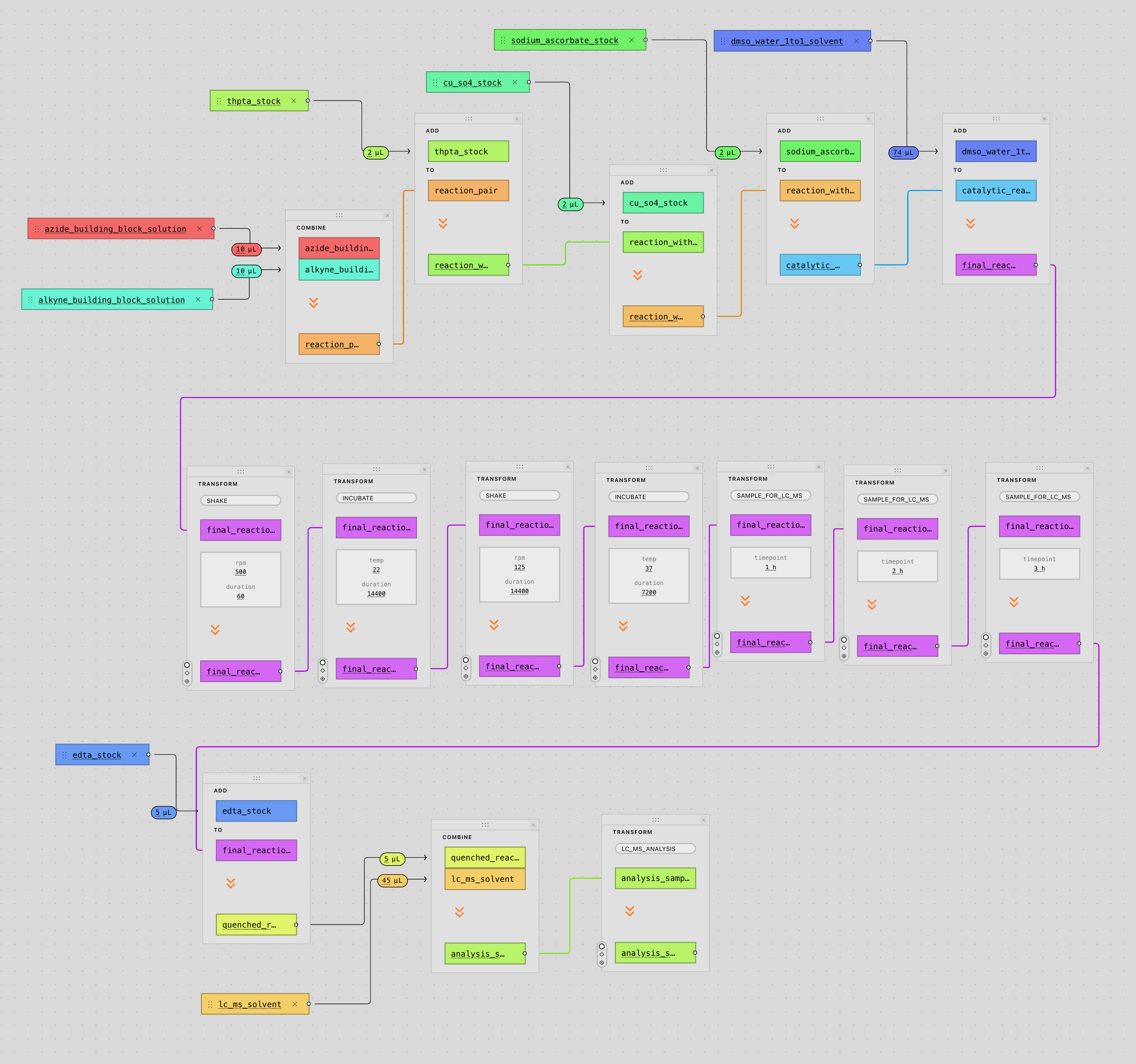
Our automation engineer, Leon, first developed a Claude skill for use in ResearchOS. Claude, when prompted with an experimental question or request, will review literature, create an experimental protocol, and use ResearchOS to control connected devices (in this case, a Hamilton STARlet, although we also have another liquid handler, an 6DoF arm, plate reader, and a few other devices connected). In this example, Leon describes the composition of a library he’d like synthesized via click chemistry, and asks Claude to generate and run the protocol. Claude then uses the ToolUniverse MCP (which provides it with access to a broad range of research tools, including BindingDB, ChEMBL, PubChem, and SwissDock) to help it respond to the prompt below before generating three artifacts it uses to get the STARlet moving with the help of ResearchOS.
Design a focused combinatorial library targeting selective inhibition of Histone Deacetylase 6 (HDAC6) via copper-catalyzed azide–alkyne cycloaddition (CuAAC) click chemistry. Identify 8 azide building blocks and 8 alkyne building blocks whose pairwise conjugation yields a structurally diverse set of candidate HDAC6-selective inhibitors. Leverage the ToolUniverse MCP to discover and employ the most relevant computational tools available for compound selection and library design.Upon completion of the literature review and library design, generate two CSV worklists formatted for robotic liquid handling, each with the headers: Well, Compound, Vol — one for the azide panel and one for the alkyne panel. Populate the Compound field with the IUPAC name of each building block.In parallel, using a separate subagent, retrieve a concise and well-documented CuAAC click chemistry protocol in PDF format suitable for upload into the Ariadne automation platform.Once all three files (azide worklist, alkyne worklist, and protocol PDF) have been prepared, invoke the Ariadne automation skill to compile the protocol and worklists into a robotic-executable experiment through the full pipeline interface. For file upload purposes, the local working directory is located at /Users/plhemstreet/Documents/ComputerUseDemo.Of course, computer use is far from the most efficient way to give a model access to ResearchOS- we would actually do this by just giving an agent access to our API. However, stopping at just giving an agent an API would be stopping short- current tools lack the transparency and standards for reproducibility that any serious tool must have, so we wanted to visualize the ability for Claude to “cowork” (so that a researcher can visually see and edit what decisions are being made) with the STARlet and researcher, as opposed to controlling it without supervision.
Many of the lessons regarding agentic experimentation we have learned in the past two years are enumerated above and in previous blogs. Our focus is far less on agentic experimentation, and more about its necessary prerequisites. Despite some potentially interesting use cases, computer use is painfully slow and token inefficient- standing up APIs/MCP servers that agents can utilize is much easier and more effective. However, there needs to be a better interface for the human user than what the terminal affords. Experimental workflows are complex and the assumptions that we, or any other tool, are making should be presented to the user in an interpretable and easily editable manner. This is a challenging UI/UX design problem. We have iterated extensively in collaboration with our users and are excited to continually evolve with them and see where it takes us.
The Path Forward
There is only one question that matters. How do we generate the highest-quality, most reproducible data at the lowest price possible? The emerging field of autonomous science contends that the answer to this question lies at the intersection of lab automation and artificial intelligence. It has been plagued by an age-old problem in the niche space of lab automation, but one that is eerily reminiscent of recurring intent specification problems of computer science. Now that we have made significant progress on this problem, we can fit the puzzle pieces of autonomous science together a lot easier.
To be frank, while we find agentic experimentation demos like this to be cool, we would not want it to draw attention away from an important fact: the challenges to the development of autonomous systems for research are not flashy. They often involve non-obvious problems with solutions that look boring next to cool hardware or models charged with developing their own novel hypotheses. It is important that as a community, we look past hype, and continue sober-minded dialogue with those who have been automating labs for decades. They have many lessons for us.
While the solutions are often not flashy or glorious in the development of this tooling, the rewards are. Our dream is basically to have a big shared research lab anyone in the world can work on, and collaborate through via this shared language/standard. We want the process of experimentation to look like the process of programming, and that requires a proper syntax and centralized/”cloud” infrastructure, in that order, just like with Oracle & SQL, and AWS & EC2. The dream is not new, but it is closer within reach than ever. A more democratic, reproducible, and easier way of interrogating secrets from the natural world is around the corner. We can’t wait to see what it brings.
Claude Max Giveaway
What types of tools might be an indispensable part of a biologist’s tool box in the era of the cloud lab? We’ve got our hands full trying to make the cloud lab happen, but we also enjoy thinking about this question, so we are giving away three 3-month subscriptions to Claude Max to students, researchers, or hobbyists that would like to use Claude Code to prototype tools for life sciences researchers. Winners will be invited to write a summary of their explorations that we will post in a blog. This giveaway was inspired by Sebastian Cocioba’s plasmid viewer, and a post that he was “treating himself” to a month of Max. People who want to build tools for biologists shouldn’t have to spend their own money on the resources to do this. While we can’t fix this problem in its entirety, we can do this. Apply here, the deadline is in 2 weeks on April 29th!
Footnotes
(1): I am not saying that anyone should actually be using LLMs for this specific purpose, just that generalized tools are getting more capable in terms of scientific reasoning across multiple modalities of input. A more capable tool would call on the specific tools that already exist (like the one developed by Kaabouch et al.), rather than just trying to turn everything into a nail because you have a hammer. Knowing how/when to use tools is important to our understanding of how to develop a better toolshed.
(2): The second summer job I had was when I was 17, trying to train a neural network to classify natural language database queries and use DeepPavlov to tag entities, so that I could turn texts coming in from Twilio into SQL commands. I wouldn’t come back to thinking deeply about NLP until roughly this moment, so it was shocking to see how much had happened in such a short amount of time. This may seem dramatic or like something we take for granted in retrospect, but it is difficult to overstate how surprising this was at the time, and how fast models have improved since.
(3): Congratulations to Will Serber, Jose Cortez, Jason Kelly and the rest of the Gingko team on this collaboration! Gingko has done wonders in bringing attention to this space with their work and has also worked to foster a very collaborative and great sense of community within the fledgling space of autonomous science. Their team has been generous and kind in their advice and help to us, as well as many others.
(4): This is not a hardware problem. We have myriad ways to pipette things and myriad ways to transfer plates between instruments. Liquid handling hardware has also gotten significantly cheaper since the Opentrons OT-2, with even Hamilton offering its ~16k Microprep. What has not changed is that putting experimental workflows onto automated platforms is still a huge pain in the ass. OwlPosting covers the different philosophies on the problems that lab automation faces in a great essay here.
(5): This is not all automation engineers do (driver integration, hardware calibration, LIMS/ELN integration, etc.) but this process of translation belongs to the set of things that they are responsible for.
(6): Much to the surprise of those outside of automation, automation engineers spend a significant amount of time understanding the biological context of the workflows they automate (this often includes attending lab meetings, reading literature, and asking researchers LOTS of questions) and very importantly, validating workflows (with dry runs, water runs, tartrazine runs).
