What Does It Mean to Be Automated?
As trite as it seems, we must clarify what we mean by “automated”. It’s a familiar word and we use it often, but we rarely explicitly define it in conversation with biologists. Sometimes we use it almost as shorthand for saying a liquid handler was used, other times we use it practically as a synonym for any kind of lab tooling at all. These definitions matter if we are to have a productive conversation about progress in lab automation.
A resource that has helped me with this question as I’ve tried to express the ideas I’ve been thinking about over the last couple years is the seminal writings on automation published by the Toyota Production System. In their framework they define and contrast two separate terms:
jidōka 自動化
(self + move + -ization)
and
jidōka 自働化
(self + human-work + -ization)
The difference is the second character 働 (hataraku) specifically implies human-like work rather than mechanical operation.
The first of these is what most likely initially comes to mind for the average English speaker upon hearing “automation” – factories are automated and robots build things on assembly lines because the work is laborious and repetitive. This is the automation that works well with the low-mix high-vol lab work that our industry has been successfully automating for years. The process engineering is tractable because the goals are clearly defined and the parameters are enumerable. If it were up to me I would replace this term with “mechanization”.
The second jidōka is sometimes translated as “automation with human touch” or “autonomation”. This is a machine that without user control will reactively make an intelligent decision. It can sense, corrects errors, and expose higher-level abstractions for the user to interface with. It reduces the burden on a human not through predefinition but intelligent feedback control.
These two kinds of automation share the same linguistic space and the same goal of reducing the need for human input, but the mindset could not be more different. Where mechanization focuses solely on achieving walk away time, autonomation is about extrapolating human intent into machine execution. Yet still, there is a third dimension of automated tooling. While mechanization dictates walk-away time and autonomation dictates the degree of abstraction, the most basic dimension of a tool is its versatility. What can it do?
What Does It Mean to Be Flexible?
Tools fall on a spectrum of versatility to specialization. Tools like liquid handlers and pipettes can be used across a much wider diversity of tasks compared to a single-function tool like a plate depeeler or sealer. In between those tools fall PCR machines and plate readers, which are capable of a neighborhood of similar tasks. The path towards improvement with specialized machines has always been straightforward. To improve it, you add intelligence – add more smart features and handle more errors and edge cases automatically. We’ve always excelled at this in biotech tooling – hell, I worked in a lab that had a qPCR machine that could connect to Alexa.
Yet, improving versatile tools without destroying their versatility is less obvious. A common approach to make a liquid handler more capable is as follows:
- The standalone liquid handler is highly versatile within its mechanical limits. It may sit idle due to usability and programming complexity, but it can perform any pattern of liquid transfers the hardware allows.
- Engineers design some number of predefined protocols the biologist runs regularly and the liquid handler can run them. Each method might get a handful of parameters and these are configurable to the scientist in a more friendly format like a worklist. Autonomy increases, and all of the process engineering decisions and individual transfers will be handled for them, but versatility decreases. I can still program my liquid handler however I please, but the interface exposes only the workflows anticipated by the engineers. Anything outside that space requires new protocol development.
- Maximum autonomy is achieved when the liquid handler is embedded into a fully integrated workcell and specialized instruments are added as modules, robotic arms move plates between instruments, and scheduling software orchestrates everything. The system runs continuously and reliably, but it only does one thing. A million-dollar biopanning workcell is useless for anything except biopanning.
Each step solves problems but we end up losing versatility and creating systems that are economically unviable for most labs. The process engineering work done to build these behemoths is remarkable and interesting and important work. However, we have taken a tool arguably more versatile than the micropipette and sacrificed its versatility for the sake of the walk-away-time of one task. It enables research at scale but it will never enable a field of high mix autonomous science.
While, autonomation reduces configuration effort required, it does not inherently reduce versatility. Adding sensors to a qPCR machine to dynamically adjust temperatures does not limit the kinds of PCR you can run. Autofocus and adaptive exposure in confocal microscopy do not reduce the breadth of imaging tasks the system is capable of. Moving from assembly language to C or Python does not restrict the computations a computer can perform. These changes reduce user burden while preserving, or even expanding, the task space.
In lab automation we routinely choose mechanization over autonomation. We call both an automated cell counter and a fully autonomous workcell “automated”, but the cell counter retains its generality with gains in ease of use, while the workcell, by contrast, often becomes ossified in its function.
Once versatility is lost it’s extremely difficult to recover. On the other hand, making something that is already versatile more intelligent is comparatively easy. Interfaces can evolve incrementally, abstractions can grow with users, and autonomy can increase without collapsing the task space. Yet, the industry has consistently conflated more capability with more mechanization.
In an effort to be more capable we adopt the wrong trajectory adapting our existing tooling. We never solved the interface of the liquid handler so we just hid behind interfaces for monolithic systems. This is inverting the abstraction hierarchy. Instead of building versatile usable primitives that compose into complex workflows, it builds complex systems that resist decomposition. Each workcell is unable to share its capabilities or the abstractions built with others and instead everything must be rebuilt for each configuration, each protocol, each variation, and for each organization.
Frameworks for Lab Automation
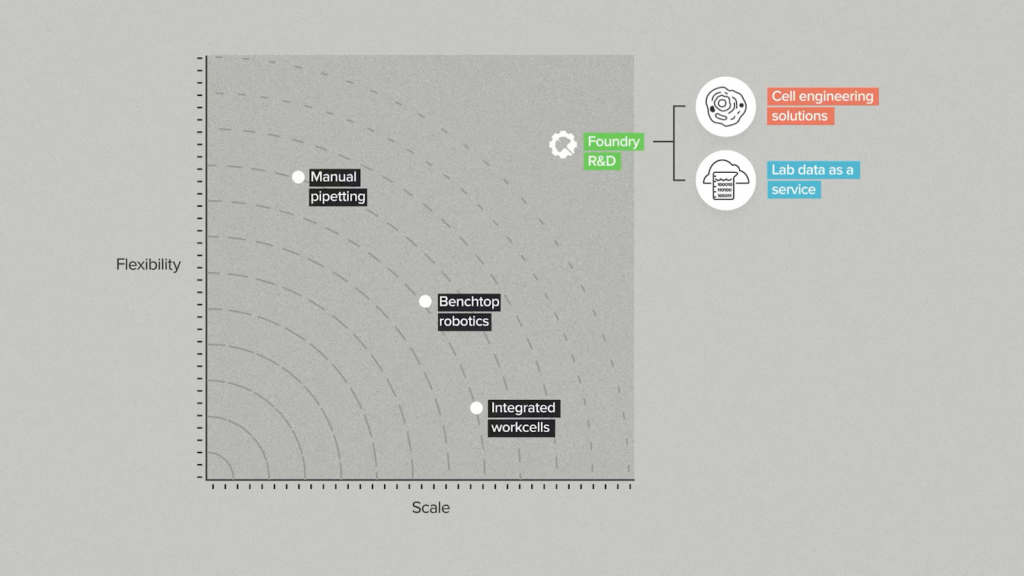
These are the terms we have been thinking about lab automation in: autonomation and versatility. I’ve noticed two similar frameworks circulating to motivate conversation around high-mix lab automation. They both classify biotech tooling along two axes. The first on the left is from Theresa Mayer, CMU’s VP of Research and the other is from Ginkgo Biowork’s Foundry series. When I initially saw these graphics I thought they might be very similar to my autonomation and versatility framing.
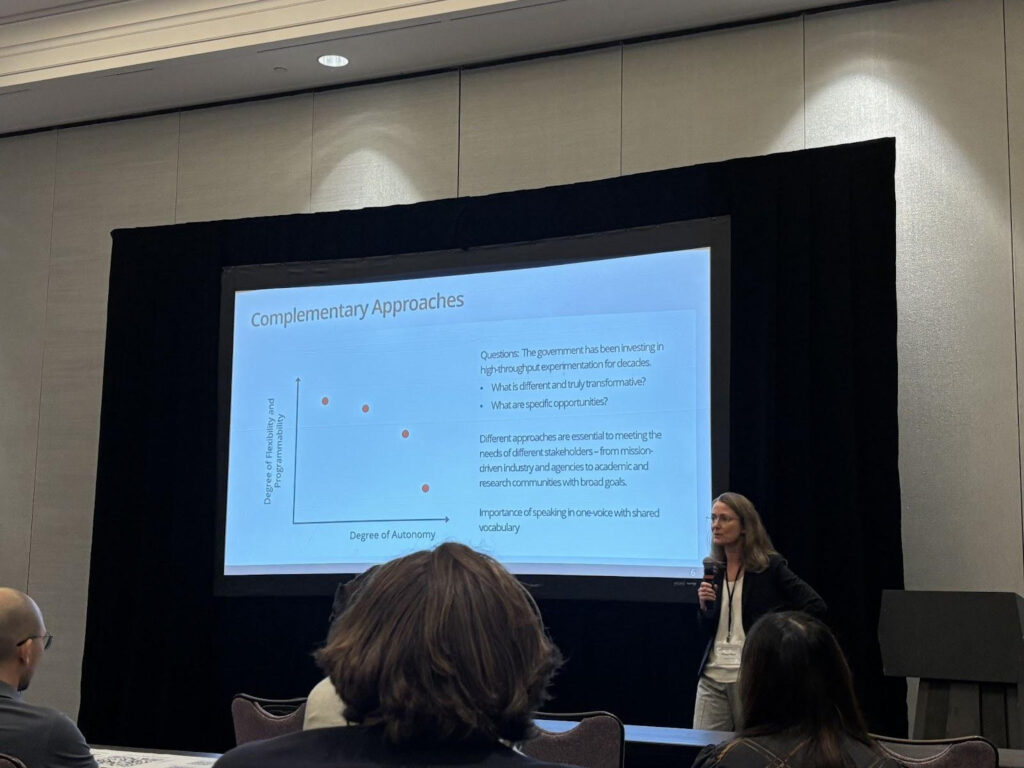
In Mayer’s slide, flexibility is used as a synonym for programmability. “Flexibility” as used here asks the question, “Given a task space, how configurable is a tool”? How much control does the user have to change the implementation of a given goal? The more ‘programmed’ something already is, the less ‘programmable’ it is and so autonomy naturally exists in opposition to this flexibility which explains her conclusion that these two dimensions must be a tradeoff between different kinds of users. The programmed kind of automation that would be low in Mayer’s flexibility is just mechanization.
This dual-axis paradigm is less powerful than autonomation and versatility because it presents a false dichotomy between automation and flexibility. This framing risks limiting thinking to solutions that apply further mechanization to low-mix/high-volume work, and more programmability for high-mix, low-volume work. If anything, that two pronged strategy for different users reflects our current practices, and what we see is that high programmability means too high an activation barrier for high-mix research labs to use automation – sure I can configure it to my needs, but now I have to invest time in configuration.
In Gingko’s framework, flexibility is used to mean what I define as versatility. Instead, effectively describing the size of the tool’s task space itself. A manual pipette isn’t flexible through “programmability”, it is flexible because it can be used for anything.
To explain why I think scale is a suboptimal choice for an axis, I’d like to borrow the software concept of horizontal scaling and vertical scaling. If you need more compute your options are to add more machines or make your machine capable of processing more at once. These two strategies have trade-offs, but that’s irrelevant to the point I am trying to make. In biology it should be an indicator that vertical scaling (each tool being capable of more scale) isn’t the bottleneck if there isn’t an associated demand for horizontal scaling (achieving scale through more machines).
Scale is a function of resources available. Robots may only be capable of a certain throughput, but the problem of augmenting scale can be solved by augmenting the number of robots. Lab automation does not suffer from a scale problem. However, greater levels of autonomation or versatility cannot be purchased – they are inherent properties of the tool.
A multi-channel isn’t simply useful because it can do 8 transfers at once, it’s useful because it abstracts the intent of the transfer from ‘transfer to these specific 8 wells’ to ‘transfers to this column’, likewise a liquid handler can abstract to an interface for ‘run a NGS workflow’. The useful dimension isn’t the scale of how many actions I can perform at once, it’s the necessary amount of human input to be specified for work. The reason why I only run one PCR a day is not because I don’t have two machines. How many biological machines spend most their time sitting idle – not least of which are liquid handlers?
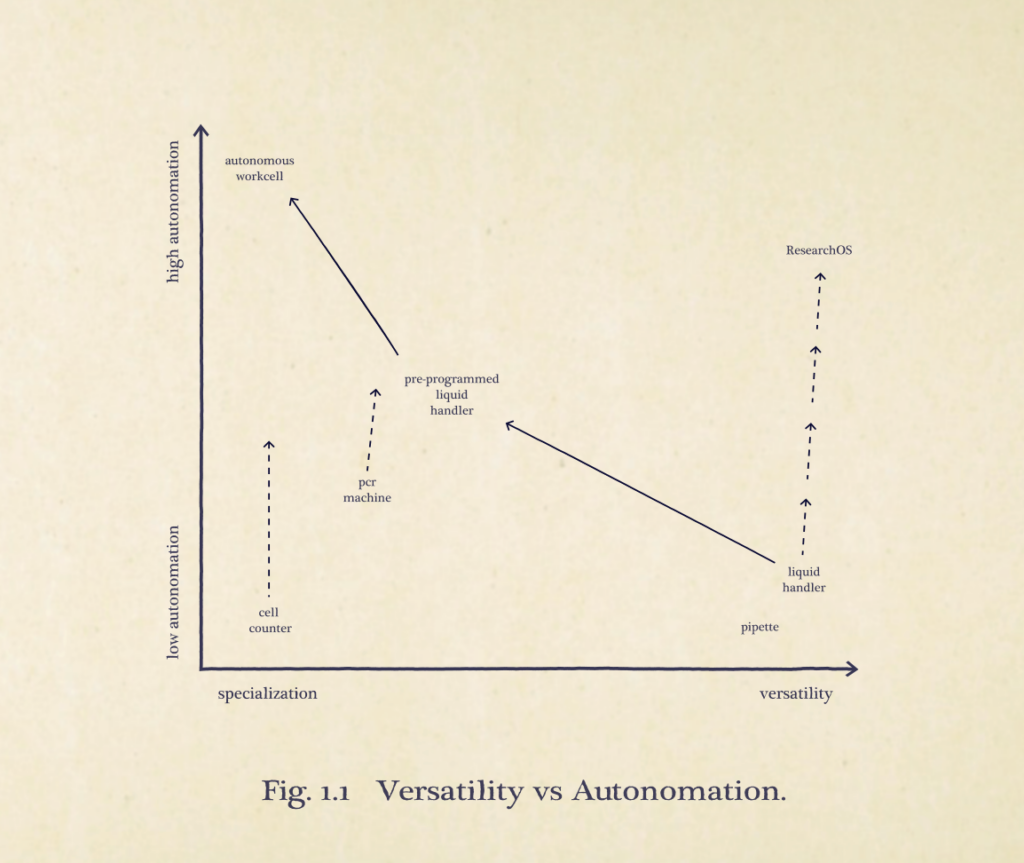
With these considerations, I present my framework. Our industry should only be working towards the paths with positive slopes. I believe that we can have more productive conversations about our goals in lab tooling. The goal is not to make every tool versatile and the goal is not to make every tool ‘fully autonomous’ running the same steps continuously forever with no input at all. The goal must be to learn how the biologist reasons, to learn what the biologist cares about, and to meet them where they are at. The only task space a scientist should have to reason in is that of their own domain, not the automation engineer’s. How they generate and evaluate automated procedures should be in their own language.
This is how high-mix automation is achieved.
P.S. I am under no delusions that the term ‘autonomation’ is going to catch on anytime soon, but the sad reality is that we don’t have much better words to delineate this concept. One could imagine an alternate history where language evolved differently in English as well and we were left with two words like in Japanese – something with roots closer to self+move+-ization and self+human-work+-ization. Historically, robotics has had “intelligent” or “agentic”, but these terms have been co-opted by genAI now, rendering them poor fits. Words that might be better fits are adaptive, responsive, mediated, autarkic, autotelic, and intentional. Ideally, I’d like something that balances both real-time sensing and the interface abstraction. Let me know what you think at rolfness at tetsuwan dot com!